Alu PCR Lab
Objective~ To determine the frequency of the Alu insert among 5 bio/biotech students.
Purpose:
Be able to list and explain the importance of each component of PCR, compare PCR to cellular DNS replication, and to associate the temperature changes with the cycling steps of PCR.
Hypothesis: There are three different genome types: minus minus (-/-), plus plus (+/+), and plus minus (+,-). We were to use our DNA to find out what genome type we were, there would be a variety of different genome types within our classroom.
Materials:
0.9% saline solution
student cheek cells
5% chelex
primer mix
master mix (nucleotides, DNA, polymerase, buffer)
pipet
paper cups
sharpie
micro-centrifuge
chelex
test tubes
agrose gel
gel box
TBE buffer
Procedure:
First we mixed our cheek cells with the saline soulution to extract some of our DNA. Then we pipetted just a little bit of it and put it in another test tube and then we centrifuged it for one minute. We then pored out the supernatant leaving the cell pellets on the bottom. After that we added chelex to our cell pellets and heated the mixture in a heat box. Once we toke it out of the heat box we centrifuged it again for one minute, then we extracted the DNA and put it in another test tube. We placed those test tubes in a rack and they were cooled overnight. The next day we pipetted some master mix and primer mix into our test tubes. We then pipetted our DNA out of the mixture and put it into another test tube. Next we made the agrose gel and put it into the gel box to cool, then we placed that in a bigger box with water. Finally we pippeted our DNA into the little slots in the gel. The next day we analyzed our gel to see how far our DNA had traveled through the gel, depending on how far it went we could tell our genome type.
Results:
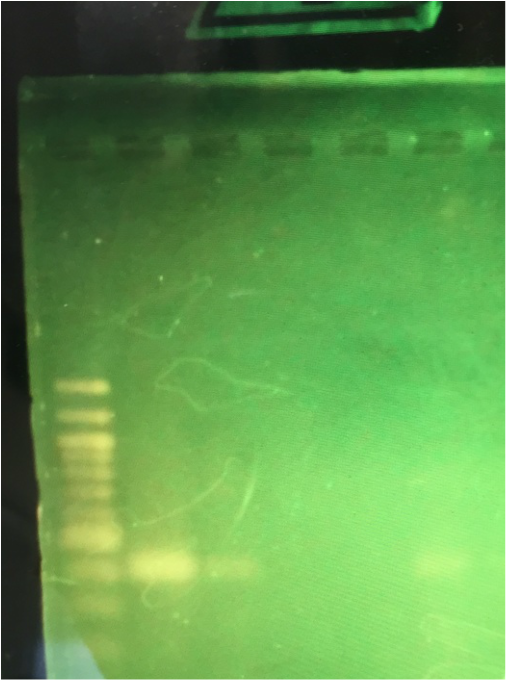
When we analyzed our gel I saw that my DNA had traveled to 400 on the base pair ladder, which means that I am a negative, negative (-,-) or a homozygus. Luckily my results were pretty bright so the band was very discernible.
Analysis:
Since my DNA turned out to be a negative negative it migrated up the gel pretty fast. Since it migrated up the gel fast that means that it was smaller in size and it didn't get caught in the gel, so it didn't get slowed down. Other people who's turned out to be positive positive meant that their DNA didn't migrate as fast as the negative negative.
Conclusion:
When I heard that we would be observing our DNA for our first lab, I was pretty excited and very intrigued on how we would do it. It was fun to put my newly learned pippetting skills it the test, and they worked. I'm very glad that my band came out bright so I was able to see my results I learned many different things during this lab, such as making sure to replace the pipet tip each time I used it so I wouldn't cross contaminate anything. Also I never knew there were so many steps in finding out someones DNA and it was really cool to get to experience how to do that step by step. I felt like there were some problems, a lot of the steps were very confusing and it was hard for the teacher to come around and help every single person. Other than that it was a very interesting lab and I learned a lot.
Purpose:
Be able to list and explain the importance of each component of PCR, compare PCR to cellular DNS replication, and to associate the temperature changes with the cycling steps of PCR.
Hypothesis: There are three different genome types: minus minus (-/-), plus plus (+/+), and plus minus (+,-). We were to use our DNA to find out what genome type we were, there would be a variety of different genome types within our classroom.
Materials:
0.9% saline solution
student cheek cells
5% chelex
primer mix
master mix (nucleotides, DNA, polymerase, buffer)
pipet
paper cups
sharpie
micro-centrifuge
chelex
test tubes
agrose gel
gel box
TBE buffer
Procedure:
First we mixed our cheek cells with the saline soulution to extract some of our DNA. Then we pipetted just a little bit of it and put it in another test tube and then we centrifuged it for one minute. We then pored out the supernatant leaving the cell pellets on the bottom. After that we added chelex to our cell pellets and heated the mixture in a heat box. Once we toke it out of the heat box we centrifuged it again for one minute, then we extracted the DNA and put it in another test tube. We placed those test tubes in a rack and they were cooled overnight. The next day we pipetted some master mix and primer mix into our test tubes. We then pipetted our DNA out of the mixture and put it into another test tube. Next we made the agrose gel and put it into the gel box to cool, then we placed that in a bigger box with water. Finally we pippeted our DNA into the little slots in the gel. The next day we analyzed our gel to see how far our DNA had traveled through the gel, depending on how far it went we could tell our genome type.
Results:
When we analyzed our gel I saw that my DNA had traveled to 400 on the base pair ladder, which means that I am a negative, negative (-,-) or a homozygus. Luckily my results were pretty bright so the band was very discernible.
Analysis:
Since my DNA turned out to be a negative negative it migrated up the gel pretty fast. Since it migrated up the gel fast that means that it was smaller in size and it didn't get caught in the gel, so it didn't get slowed down. Other people who's turned out to be positive positive meant that their DNA didn't migrate as fast as the negative negative.
Conclusion:
When I heard that we would be observing our DNA for our first lab, I was pretty excited and very intrigued on how we would do it. It was fun to put my newly learned pippetting skills it the test, and they worked. I'm very glad that my band came out bright so I was able to see my results I learned many different things during this lab, such as making sure to replace the pipet tip each time I used it so I wouldn't cross contaminate anything. Also I never knew there were so many steps in finding out someones DNA and it was really cool to get to experience how to do that step by step. I felt like there were some problems, a lot of the steps were very confusing and it was hard for the teacher to come around and help every single person. Other than that it was a very interesting lab and I learned a lot.
